Publications
The citations on this page were generated automatically using the Manubot cite utility developed in the Greene Lab, which also developed the template for this website.
Featured

Long-read metagenomics reveals phage dynamics in the human gut microbiome
Nature
·
26 Nov 2025
·
doi:10.1038/s41586-025-09786-2
For this paper, we aimed to study prophages in the human gut microbiome using long-read sequencing. How stable are prophages in their bacterial hosts? How broad are prophage host ranges? What type of integration strategies do prophages use in the gut? Read more if you also want to learn about a new class of ‘IScream phages’ that we discovered.
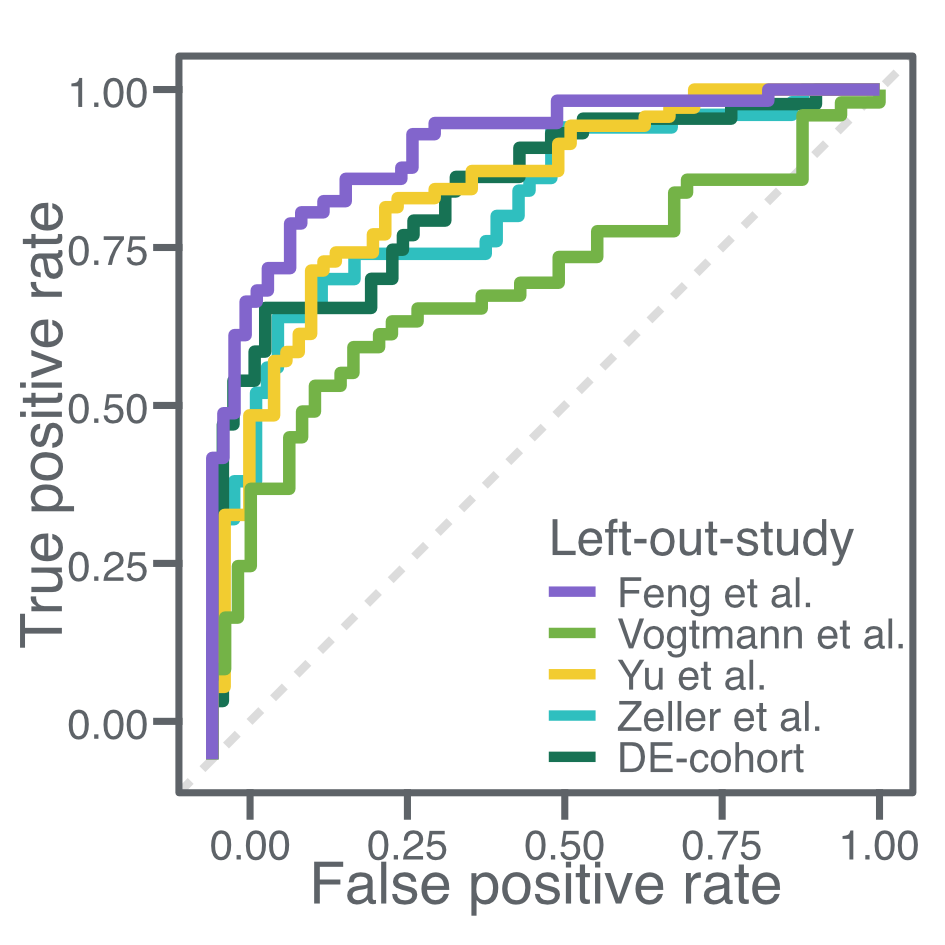
Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer
Nature Medicine
·
01 Apr 2019
·
doi:10.1038/s41591-019-0406-6
In this study, we asked if microbial biomarkers for colorectal cancer are generalizable across different datasets, populations, and geographies. We found a set of bacterial species that were highly enriched in colorectal cancer metagenomes and could train an accurate and specific machine learning model for the detection of colorectal cancer.
All
2026
Differential effects of two common GVHD prophylaxis regimens on the gut microbiome: Results from the BMT CTN 1801 study
openRxiv
·
20 Feb 2026
·
doi:10.64898/2026.02.19.706769
2025
Long-read metagenomics reveals phage dynamics in the human gut microbiome
Nature
·
26 Nov 2025
·
doi:10.1038/s41586-025-09786-2
A peptide display system identifies a potent mutant β-melanocyte-stimulating hormone agonist of melanocortin-4 receptor
Cell Genomics
·
01 Nov 2025
·
doi:10.1016/j.xgen.2025.100988
ATF6 activation alters colonic lipid metabolism causing tumour-associated microbial adaptation
Nature Metabolism
·
01 Sep 2025
·
doi:10.1038/s42255-025-01350-6
The prototypic crAssphage is a linear phage-plasmid
Cell Host & Microbe
·
01 Aug 2025
·
doi:10.1016/j.chom.2025.07.004
Machine learning-based meta-analysis reveals gut microbiome alterations associated with Parkinson’s disease
Nature Communications
·
07 May 2025
·
doi:10.1038/s41467-025-56829-3
Accurate prediction of absolute prokaryotic abundance from DNA concentration
Cell Reports Methods
·
01 May 2025
·
doi:10.1016/j.crmeth.2025.101030
Expanding the human gut microbiome atlas of Africa
Nature
·
29 Jan 2025
·
doi:10.1038/s41586-024-08485-8
2024
Intragenic DNA inversions expand bacterial coding capacity
Nature
·
25 Sep 2024
·
doi:10.1038/s41586-024-07970-4
A realistic benchmark for differential abundance testing and confounder adjustment in human microbiome studies
Genome Biology
·
25 Sep 2024
·
doi:10.1186/s13059-024-03390-9
2023
C. difficile may be overdiagnosed in adults and is a prevalent commensal in infants
eLife Sciences Publications, Ltd
·
12 Sep 2023
·
doi:10.7554/eLife.90111.1
Location and condition based reconstruction of colon cancer microbiome from human RNA sequencing data
Genome Medicine
·
02 May 2023
·
doi:10.1186/s13073-023-01180-9
2022
Cultivation-independent genomes greatly expand taxonomic-profiling capabilities of mOTUs across various environments
Microbiome
·
05 Dec 2022
·
doi:10.1186/s40168-022-01410-z
Depression and fatigue in active IBD from a microbiome perspective—a Bayesian approach to faecal metagenomics
BMC Medicine
·
17 Oct 2022
·
doi:10.1186/s12916-022-02550-7
Microbiota-dependent activation of the myeloid calcineurin-NFAT pathway inhibits B7H3- and B7H4-dependent anti-tumor immunity in colorectal cancer
Immunity
·
01 Apr 2022
·
doi:10.1016/j.immuni.2022.03.008
Calorie restriction improves metabolic state independently of gut microbiome composition: a randomized dietary intervention trial
Genome Medicine
·
14 Mar 2022
·
doi:10.1186/s13073-022-01030-0
A faecal microbiota signature with high specificity for pancreatic cancer
Gut
·
08 Mar 2022
·
doi:10.1136/gutjnl-2021-324755
Identifying temporal and spatial patterns of variation from multimodal data using MEFISTO
Nature Methods
·
13 Jan 2022
·
doi:10.1038/s41592-021-01343-9
2021
Unravelling the collateral damage of antibiotics on gut bacteria
Nature
·
13 Oct 2021
·
doi:10.1038/s41586-021-03986-2
Commensal Clostridiales strains mediate effective anti-cancer immune response against solid tumors
Cell Host & Microbe
·
01 Oct 2021
·
doi:10.1016/j.chom.2021.08.001
Prediction of combination therapies based on topological modeling of the immune signaling network in multiple sclerosis
Genome Medicine
·
16 Jul 2021
·
doi:10.1186/s13073-021-00925-8
Microbiome meta-analysis and cross-disease comparison enabled by the SIAMCAT machine learning toolbox
Genome Biology
·
30 Mar 2021
·
doi:10.1186/s13059-021-02306-1
2020
Metabolic models predict bacterial passengers in colorectal cancer
Cancer & Metabolism
·
10 Feb 2020
·
doi:10.1186/s40170-020-0208-9
2019
Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation
Nature Medicine
·
01 Apr 2019
·
doi:10.1038/s41591-019-0405-7
Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer
Nature Medicine
·
01 Apr 2019
·
doi:10.1038/s41591-019-0406-6
Extensive transmission of microbes along the gastrointestinal tract
eLife
·
12 Feb 2019
·
doi:10.7554/eLife.42693
2018
Phosphoproteomics-Based Profiling of Kinase Activities in Cancer Cells
Methods in Molecular Biology
·
01 Jan 2018
·
doi:10.1007/978-1-4939-7493-1_6