Software

SIAMCAT
The SIAMCAT package is an R package for comparative metagenomics analysis. It is a one-stop toolbox for data filtering, association testing, construction of machine learning workflows, and data visualization.
Check out the associated publication describing the tool.
The SIAMCAT package was originally developed in the Zeller lab.
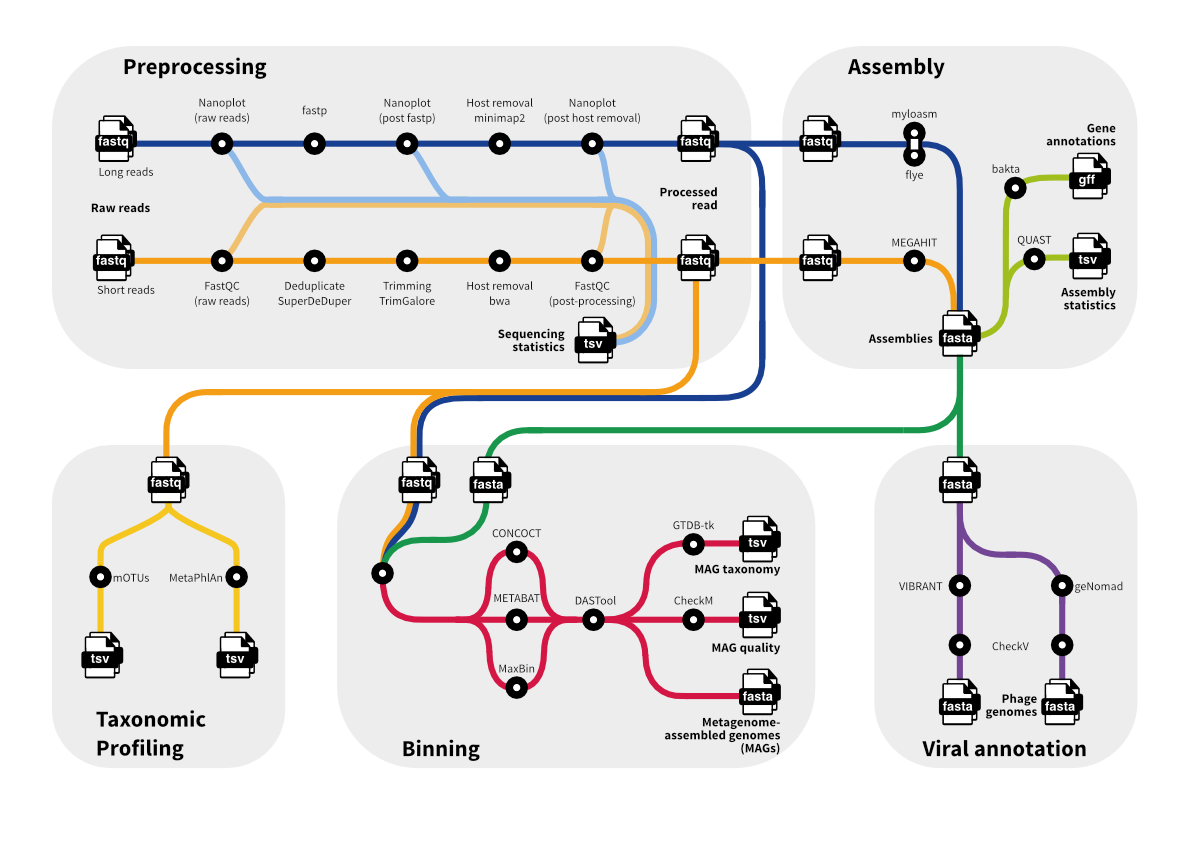
Microbiome Workflows
The microbiome workflows are implemented in Nextflow and offer a wide range of standard microbiome analyses approaches. They include workflows for raw data preprocessing, taxonomic profiling, assembly, binning, and prediction of phages. Most workflows are implemented to work with both short- and long-read input data.
The microbiome workflows are maintained by the Bhatt lab and the Wirbel lab.